What we do..
We aim to discover novel fundamental biology in the areas of Immunometabolism, Cancer Immunology and Aging by using cross-disciplinary approaches to data generation and analysis together with in vitro and in vivo experimental biology.In our laboratory, classically trained immunologists and molecular biologists work hand-in-hand with computational biologists and computer scientists to leverage the power of high-throughput omics datasets.
Collectively, our approach can be characterized as Systems Immunology.
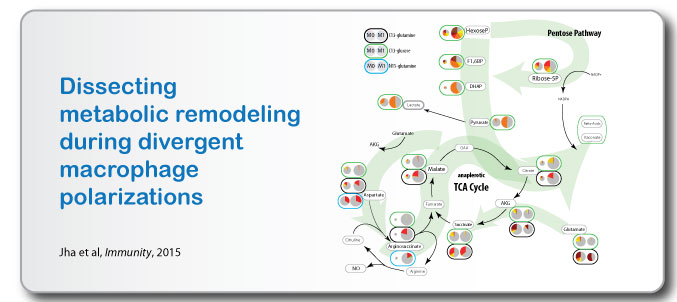
- Immunometabolism One major example of this approach is our dissection of the metabolic rewiring of M1- and M2-polarized macrophages (Jha et al, 2015) that highlighted metabolic breakpoints of TCA cycle and the redirection of the metabolic flux to itaconate in M1 macrophages, as well as glutamine and UDP-GlcNAc dependence of M2 polarized macrophages.
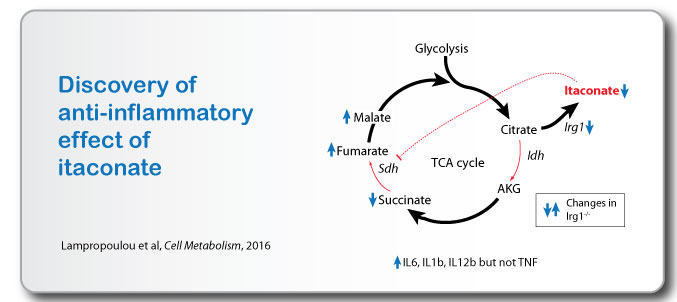
This systematic work led us to discovery of anti-inflammatory action of itaconate (Lampropoulou et al, 2016) where using mice lacking endogenous itaconate (Irg1-/-) we show that itaconate inhibits Sdh and inhibits production of pro-inflammatory cytokines Il1b, Il6 and Il12b.
In our most recent work we show that itaconate is also a natural electrophile (Bambouskova et al, 2018) and can trigger electrophilic stress response when produced in activating macrophages: we show that induces both Nrf2 and Atf3 driven responses. Importantly, this property of itaconate can be chemically enhanced by using certain itaconate derivatives to treat autoimmune diseases associated with Ikbz disfunction, such as psoriasis.